Selected publications and presentations
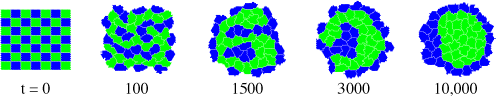
Minsuk, S. B. (2005). Toward an open-ended and mechanically realistic model of biological cells. In: Workshop on Artificial Chemistry and Its Applications. The 8th European Conference on Artificial Life (ECAL) Workshop Procedings. Mathieu Capcarrere (ed.). [download pdf]
Minsuk, S. B. (2004). First steps toward a generalized model cell for evo-devo computer simulations. Abstract: Integrative and Comparative Biology 44:729. (A poster presented at the Society for Integrative and Comparative Biology, Jan. 2005.)
[View poster, with QuickTime movies of simulations]
Minsuk, S. (2002). A proposal for an artificial life simulation of evolving multicellular ontogeny. (A poster presented at Artificial Life VIII [8th International Conference on the Simulation and Synthesis of Living Systems], Dec. 2002.)
[abstract only]
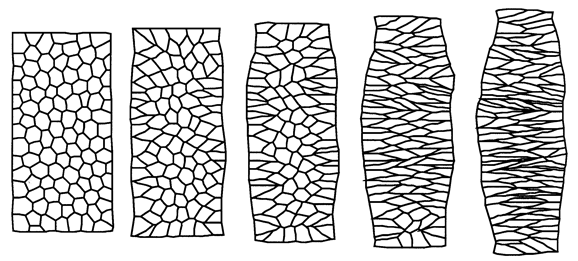
Weliky, M., S. Minsuk, R. Keller and G. Oster (1991). Notochord morphogenesis in Xenopus laevis: simulation of cell behavior underlying tissue convergence and extension. Development 113:1231-1244.
Oster, G., M. Weliky, and S. Minsuk (1990). Morphogenesis by cell intercalation. In: 1989 Lectures in Complex Systems. Santa Fe Institute Studies in the Sciences of Complexity Series, Lect. Vol. II. 501-512. Erica Jen (ed). Reading, MA:Addison-Wesley.
|